Barbora Špačková of Chalmers University of Technology, Sweden, secures the chip in a specially adapted dark-field microscope and illuminates it with visible light. [Image: M. Saaranen, Envue Technologies]
Fluorescence microscopy has many applications in biophysics, but labeling nanoparticles and biomolecules with fluorescent tags can alter the properties of the tiny bits. Scientists have devised several optical detection strategies to avoid labeling single biomolecules, but all these methods require the molecular species to bind to a surface—and not all molecules and nanoparticles can do that.
Now, researchers in Sweden have developed a new technique called nanofluidic scattering microscopy, which images the target particles while they are flowing through an optically transparent matrix of nanoscale channels (Nat. Methods, doi: 10.1038/s41592-022-01491-6). The technique, which incorporates dark-field light-scattering microscopy, makes it possible to determine molecular weights and other important data for single biomolecules in solution.
How the setup works
The researchers designed their experimental setup with a dark-field microscope, a polychromatic visible-light source and a CMOS camera running at 200 frames per second. In the center of this gear was a nanofluidic chip designed by the researchers with channels 30 to 200 μm in length and widths of a few tens or hundreds of nanometers.
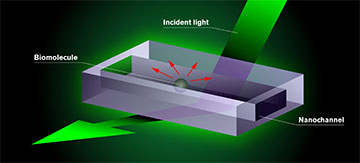
The biomolecules the researchers want to study are placed in a chip consisting of nanochannels. Test fluid is added to the chip, which is mounted in an optical dark-field microscope and illuminated with visible light. The molecule appears as a dark shadow moving freely inside the channel on a screen connected to the microscope. The darkness of the shadow is proportional to the mass of the molecule. This is the result of an interference effect in the interaction of the light with the nanochannel and the molecule. [Image: Chalmers University of Technology / Y. Strandqvist and D. Spacek, Neuron Collective] [Enlarge image]
As an inert fluid pumps the tiny particles through the nanochannels, the camera records a dark-field image of the light scattering off the particles. The researchers then generate a differential dark-field image by subtracting an image of the empty channel from the image of the channel with the particle flowing through it.
The nanochannels keep the particles of interest within the focal plane of the microscope at all times and improve the optical contrast of the particles by several orders of magnitude.
Seeing tiny molecules
According to Chalmers University of Technology physicist Christoph Langhammer, nanofluidic scattering microscopy can image individual molecules down to the size range of a few tens of kilodaltons. (A dalton is a unit of atomic mass, 1/12 of the mass of a carbon-12 atom.) The method is more efficient than other label-free microscopy schemes because scientists “can derive both a biomolecule’s mass (from optical contrast) and hydrodynamic radius (from diffusivity) from a single measurement using a single technique,” Langhammer adds.
According to Langhammer, the biggest challenge the team faced was reaching the shot-noise limit in their experimental setup. Next, the researchers will push the technique’s detection limit toward even smaller molecules by improving the design of the nanofluidic channels and refining their data analysis.