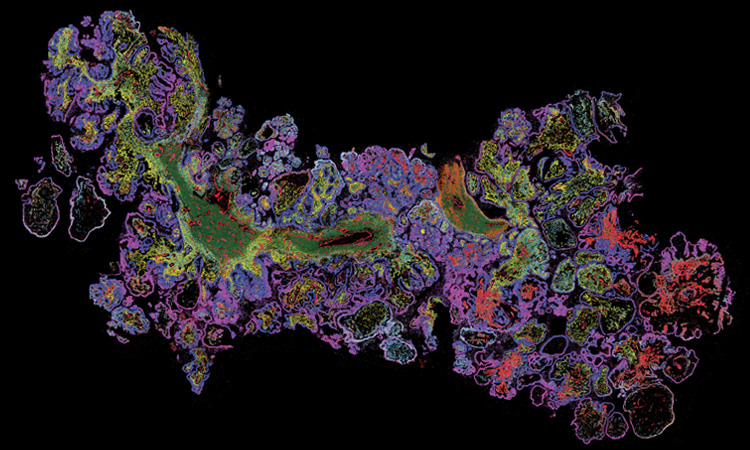
 MERSCOPE data image from a human ovarian cancer tissue, displaying nine genes in different colors. [Vizgen Inc.]
MERSCOPE data image from a human ovarian cancer tissue, displaying nine genes in different colors. [Vizgen Inc.]
In 1977, British biochemist Frederick Sanger and his colleagues sequenced the first DNA-based genome—a circular, single-stranded DNA made up of 5,386 nucleotides—from a bacteriophage that infects Escherichia coli. The demonstration showed off the ability of the “Plus and Minus” system, known as the first-ever DNA sequencing technique. Later that same year, the team introduced a second DNA sequencing method, which represented a vast improvement over all others, including its own “Plus and Minus” approach.
Now referred to as Sanger sequencing, the breakthrough forever altered the progress of DNA sequencing technology, allowing scientists to rapidly and accurately sequence long stretches of DNA with ease. The advent of Sanger sequencing opened the floodgates for a flurry of genome sequencing projects, from mitochondria and bacteria to fruit flies and zebrafish. In 2003, the Human Genome Project produced an accurate and complete human genome, comprising over 3 billion nucleotides.
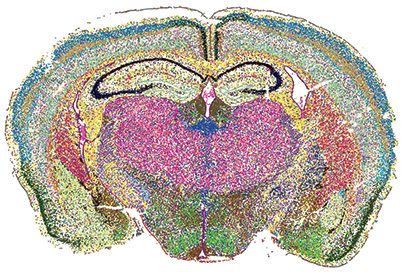 MERSCOPE PanNeuro Cell Type 500 Gene Panel data image displaying spatial distribution of identified cell types across a mouse brain coronal section. [Renchao Chen, Vizgen Inc.]
MERSCOPE PanNeuro Cell Type 500 Gene Panel data image displaying spatial distribution of identified cell types across a mouse brain coronal section. [Renchao Chen, Vizgen Inc.]
The world of omics
The omics “family” is composed of many disciplines, including the examples below:
Genomics: Study of a complete set of genes
Transcriptomics: Study of a complete set of RNA transcripts
Proteomics: Study of a complete set of expressed proteins
Epigenomics: Study of a complete set of reversible chemical modifications to DNA
Metabolomics: Study of a complete set of small-molecule metabolites
Today, it’s appreciated that genomics—which, as opposed to genetics, focuses on a whole genome rather than individual genes—makes up only one piece of the puzzle. The recent “omics” revolution in the biological sciences aims to make a comprehensive assessment of a set of molecules in a cell, tissue or organism. During the past several years, a new trend termed “spatial omics” has seen a meteoric rise in popularity. It applies omics techniques to intact tissue samples instead of dissociated cells, allowing researchers to create a biomolecular map by overlaying omics data onto tissue images. In this way, spatial omics methods are able to retain the native context of biomolecules within cells, and of cells within a larger tissue sample.
“One way to think about spatial omics as a field is the merger of two different ways that people have historically been looking at biological samples: microscopy and omics,” said Jeffrey Moffitt, assistant professor of cellular and molecular medicine at Boston Children’s Hospital, USA. “It captures all the strengths of these two very different fields and combines them together in a way that offsets their weaknesses.” Nature Methods crowned spatially resolved transcriptomics the “Method of the Year” in 2020, stating that it “stands to change the way cell biology as well as pathology and histology are practiced.”
And optics and photonics are key to exploring this new frontier—from the microscopy used to produce tissue images onto which omics data are overlaid, to the fluorescent tags needed to label hundreds of different molecules.
From bulk to single-cell sequencing
Before examining the optics behind spatial omics, it is useful to understand why these techniques have become so valuable to scientists. While Sanger sequencing could only work on one strand of DNA at a time, next-generation sequencing—initially called “massively parallel sequencing” when first introduced in the 1990s—had the power to sequence many strands simultaneously. Next-generation sequencing facilitated a new era of genomics where studies could be completed quickly and cost effectively.
In the decades that followed, scientists sequenced thousands of genomes from numerous species by extracting DNA from millions of cells and pooling them together. For all its strengths, next-generation sequencing at that time could only produce an average of many cells and was unable to analyze a small number of cells. But since the cell is the basic unit of biology, and averaging loses the heterogeneity among individual cells, researchers wanted to be able to sequence a single cell.
Single-cell sequencing leverages optimized next-generation sequencing technologies to detect the genome, transcriptome and other “omes” from single cells. Early methods were prohibitively expensive, but over time, lower-cost approaches emerged. Today, high-throughput single-cell sequencing has become more widely available to distinguish among cell types and understand cell-to-cell relationships in areas such as oncology, neurology and immunology. “Over the past 10 years, single-cell sequencing has really revolutionized the way we think about biology,” said Moffitt. “Any individual tissue might be composed of hundreds of different types of cells, and each of those cells plays a different role. And for many tissues, we just didn’t really know what cells were there.”
The technology has enabled the more accurate, rapid identification of rare and novel cells in tissue, while also providing unprecedented insight into cell composition and function. However, the process of isolating a single cell for analysis still left something to be desired—namely, the spatial context of the cell among its neighbors and surrounding environment.
“The field of single-cell sequencing is still very much alive, and people try to understand disease by examining which cell types are present in the tissue,” said Carolina Wählby, professor of quantitative microscopy at Uppsala University, Sweden. “With in situ sequencing, we can then also see where the cell types are in the tissue.”
To get a better handle on the spatial organization of cells within a tissue, researchers have pressed into service a number of optics-related techniques, especially in the fluorescence realm.
The evolution of FISH
To get a better handle on the spatial organization of cells within a tissue, researchers have pressed into service a number of optics-related techniques, especially in the fluorescence realm. Early approaches include laser capture microdissection (LCM) and single-molecule fluorescent in situ hybridization (smFISH). LCM uses either a UV or infrared laser to excise a micrometer-scale region of interest (ROI) or even single cells from a sample. From there, full gene or protein profiling at a single-cell level can be performed. But the process of dissecting cells is laborious and not ideal for mapping a whole tissue section.
In fluorescent in situ hybridization (FISH), fluorescent probes bind to a target nucleic acid sequence to detect and localize DNA or RNA in individual cells or tissues. The probe consists of a single strand of DNA or RNA, tagged with fluorophores, that is complementary to the target sequence. After taking steps to hybridize and wash away any unbound probes, fluorescence microscopy is used to visualize any bound FISH probes in the sample.
To generate a high enough signal for detection, FISH requires a sufficient concentration of the target sequence to provide contrast against surrounding areas. smFISH, developed in the late 1990s, modified FISH to allow for the detection of single RNA molecules. Instead of one long probe, the researchers made several short probes—called oligonucleotides—that would bind to adjacent sequences on a target RNA molecule. With this strategy, the collective fluorescence of a single messenger RNA (mRNA) molecule could then be detected in a wide-field fluorescence microscopy image.
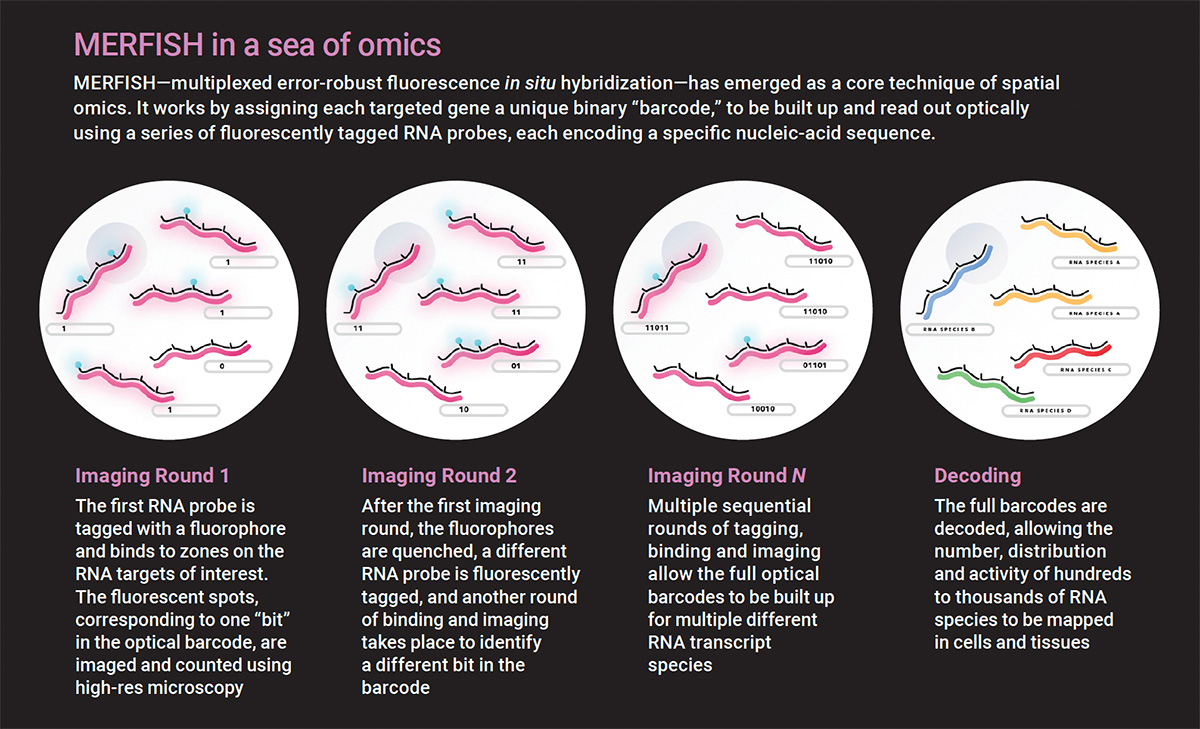
smFISH has evolved into several highly multiplexed methods, including MERFISH (multiplexed error-robust fluorescence in situ hybridization). MERFISH incorporates combinatorial labeling, sequential imaging and error-robust barcoding to encode hundreds of genes. Each targeted gene is assigned a unique binary optical barcode, and each round of imaging corresponds to one bit. Between rounds, the fluorescent tags are inactivated to make the sample dark again. Multiple sequential rounds of imaging are then performed to read out the full optical barcode.
“Each molecule will have its own fluorescent on-off pattern that you can think of as a barcode,” said Moffitt, the co-inventor of MERFISH technology. “Now you know where every single molecule was because you have these fluorescent spots, and from the barcodes, you know the identity of that molecule.”
 The Vizgen MERSCOPE platform combines single-cell and spatial transcriptomics analysis with spatial imaging for a commercial application of MERFISH technology. [Vizgen Inc.]
The Vizgen MERSCOPE platform combines single-cell and spatial transcriptomics analysis with spatial imaging for a commercial application of MERFISH technology. [Vizgen Inc.]
His laboratory is applying MERFISH to questions associated with the gastrointestinal tract, such as its cellular composition and organization, the triggers of inflammation and disease, and the response of the gut to the microbiome. Recently, Moffitt and his colleagues used MERFISH to profile the expression of 940 genes in 1.35 million cells imaged across onset and recovery in a mouse colitis model. They have also commercialized the technology in the form of the Vizgen MERSCOPE Platform.
“One of the exciting things about creating a technique like MERFISH is that almost every biological question in a tissue comes down to what are the cell types and how are they organized,” he said. “MERFISH is really a technique that can be applied very broadly across many different organisms in many different tissue contexts.”
Antibody-based imaging
Antibodies form the basis for several spatial proteomics methods, since they are able to bind to a specific protein of interest. Most commonly, antibodies are labeled with metals, fluorophores or DNA oligonucleotides that are then detected by mass spectrometry or fluorescence microscopy.
Most commonly, antibodies are labeled with metals, fluorophores or DNA oligonucleotides that are then detected by mass spectrometry or fluorescence microscopy.
For instance, imaging mass cytometry (IMC) incorporates metal-conjugated antibodies along with a UV laser that vaporizes the tissue while scanning the sample pixel by pixel. The metals released from each pixel are then quantified with mass spectrometry. IMC allows for the simultaneous detection of up to 50 antigens, with the limiting factor being the availability of metal isotopes.
Alternatively, cyclic immunofluorescence relies on the detection of fluorescent antibodies over repeated imaging cycles. High-dimensional datasets with hundreds of antigens can be obtained with an iterative, multistep process. First, antibodies are tagged with fluorophores. Next, images are acquired using diverse imaging systems, including laser scanning confocal microscopes capable of high-parameter imaging. Finally, either antibodies are removed or fluorophores are inactivated to prevent spectral overlap.
“We might apply three fluorescently conjugated antibodies per cycle, and for a 90-plex image, that means you would have to do 30 cycles of dye inactivation or antibody removal,” said Andrea Radtke, staff scientist/associate scientist at the National Institute of Allergy and Infectious Diseases (NIAID), USA.
In an effort to increase efficiency, Radtke led the development of IBEX (“iterative bleaching extends multiplexity”), a versatile multiplex optical imaging approach that can be completed in two to five days by biologists with basic laboratory skills. IBEX uses a rapid chemical bleaching agent to bleach more than 15 unique fluorophores without tissue destruction. The method significantly reduces the fluorophore inactivation step and antibody labeling time from over 16 hours to less than one hour.
 [Enlarge image][Adapted from Vizgen Inc.]
[Enlarge image][Adapted from Vizgen Inc.]
One of the most widely used cyclic immunofluorescence methods is CODEX (“codetection by indexing”), which has the advantage of binding all the antibodies to the sample at once in a single step. In CODEX, each antibody is tagged with a DNA oligonucleotide, and the complementary sequence is conjugated to a fluorescent reporter dye. Detection is performed in cycles, during which fluorescently labeled oligonucleotides are applied to the tissue three colors at a time. Images are captured using an inverted fluorescence optical microscope. Then, the fluorescent probes are removed, and the cycle repeats itself.
Radtke is also an associate member of the Human BioMolecular Atlas Program (HuBMAP), an ambitious initiative to create a multiscale spatial atlas of the healthy human body at single-cell resolution. The overarching goal of the project is to generate reference spatial maps of functional tissue units across many organs from diverse populations to help determine how the relationships among cells can affect the health of an individual.
Founded in 2018, HuBMAP involves more than 60 institutions and 400 researchers from around the world. The project, currently in the production phase, employs a laundry list of different technologies for mapping: antibody-based imaging, spatial transcriptomics, single-cell sequencing, smFISH and histological imaging, among others. “Optics is behind this big effort to create a human reference atlas,” said Radtke. “We know that there are 37 trillion cells in the human body, and there’s been a recent explosion of technologies that we can now use to catalog and map these cells.”
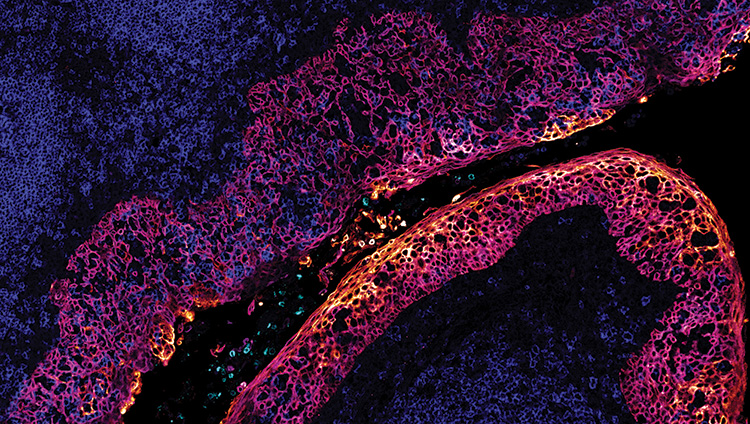 IBEX image of a human tonsil by Andrea Radtke of Ronald Germain’s lab at NIAID, USA. [NIAID]
IBEX image of a human tonsil by Andrea Radtke of Ronald Germain’s lab at NIAID, USA. [NIAID]
In situ sequencing
One challenge for spatial proteomics methods based on multiplexed antibody detection is that the antibodies themselves are imperfect tools whose behavior remains highly sensitive to sample preparation conditions. “You can look for proteins, but then you’re dependent on having good antibodies that are very specific,” said Wählby. “What we do instead is to look for mRNA and the transcriptomics—which is, of course, not the same thing as the protein, but it’s still relevant information to decide what cell types we have in different parts of the tissue.”
The tool enabling that search is in situ sequencing, a family of methods characterized by the high-throughput sequencing of mRNA directly in preserved cells or tissues. In 2013, Wählby and her colleagues developed one of the first in situ sequencing technologies based on “padlock” DNA probes hybridized to mRNA.
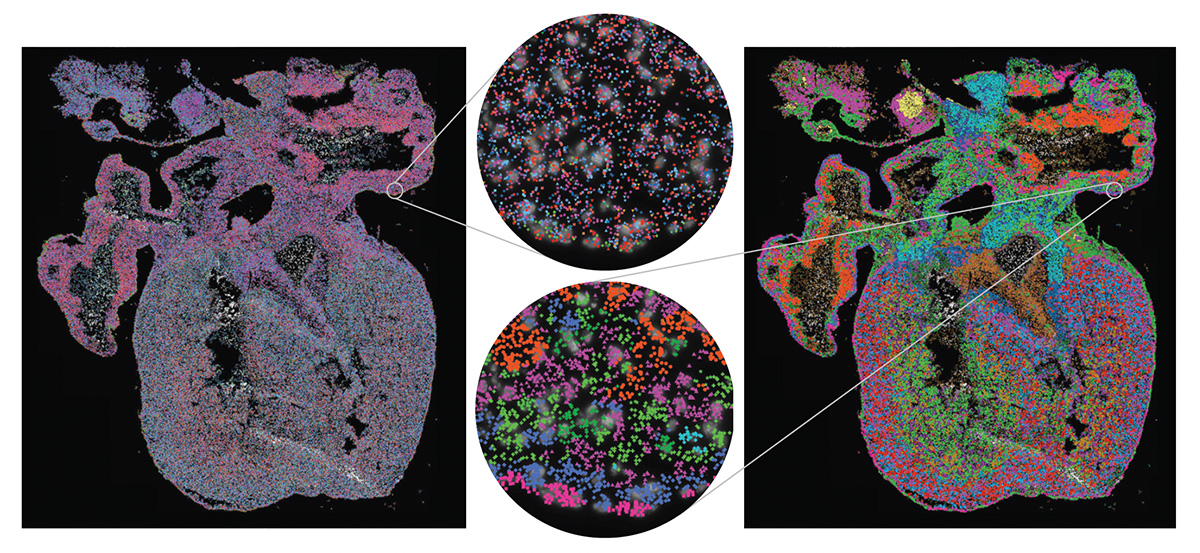 [Enlarge image]Left: Gene expression in the developing human heart. Each marker represents one detected mRNA. Right: The same data, but with expressed genes grouped into cell types to visualize the cellular architecture. [TissUUmaps / Uppsala University]
[Enlarge image]Left: Gene expression in the developing human heart. Each marker represents one detected mRNA. Right: The same data, but with expressed genes grouped into cell types to visualize the cellular architecture. [TissUUmaps / Uppsala University]
First, a DNA probe attaches to a target mRNA in the form of a circle. The circular DNA reacts with enzymes that begin going around the circle, making many copies. This isothermal process, called rolling-circle amplification, creates a large “blob” that can be visualized with a fluorescence microscope at micrometer spatial resolution. From there, the mRNA is sequenced by ligation, a technique that uses probe oligonucleotides labeled with fluorescent dyes to decode bases one by one.
“Since 2013, there have been lots of applications of in situ sequencing, as well as different tricks and tweaks to make it more efficient,” said Wählby. “There are also new ways of decoding these blobs. Initially, we used fairly simple FISH probes, but many other versions of probes have come after that to make the decoding process faster, more sensitive and more efficient.”
Recently, Wählby and her colleagues created the first topographic atlas of the developing human lung by combining gene expression profiling via single-cell RNA sequencing with in situ sequencing on intact tissues. They investigated lung tissue from 17 embryos ranging from 5 to 14 weeks post-conception, identifying and mapping 83 cell states as well as multiple developmental trajectories as cells progressed from immature to mature.
Imaging was performed using a Zeiss Axio Imager.Z2 epifluorescence microscope, with a Zeiss Plan-Apochromat 20×/0.8 objective and an automatic multislide stage to facilitate the repetitive-cycle imaging needed for sequencing by ligation. The system was equipped with a Lumencor SPECTRA X light engine LED source, having the 395/25, 438/29, 470/24, 555/28, 635/22 and 730/40 filter paddles.
“There are two different areas where I think in situ sequencing is relevant. One is development—if you understand the process of development, that’s when you can also understand or hypothesize about what happens when it goes wrong,” she said. “Then, of course, the really big application area is in tumor biology.”
Her laboratory just completed work on the interface between tumor and normal tissue during liver metastasis in patients with colon cancer. In certain cases, tumor cells are encapsulated by the liver, which makes them easier to remove surgically. She hopes that spatial omics techniques can map the interface and give us a better understanding of why this encapsulation happens in some patients but not others.
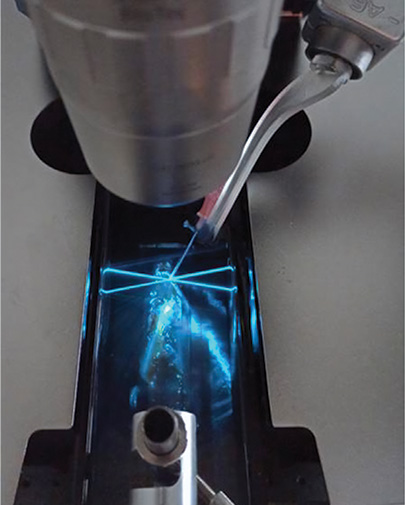 A method called DISCO-MS leverages AI and robotics to yield spatial proteomics data in optically cleared whole specimens. [Reprinted from H. Singh Bhatia et al., Cell 185, 5040 (2022), with permission from Elsevier.]
A method called DISCO-MS leverages AI and robotics to yield spatial proteomics data in optically cleared whole specimens. [Reprinted from H. Singh Bhatia et al., Cell 185, 5040 (2022), with permission from Elsevier.]
The future of spatial omics
Many other spatial profiling technologies exist that call for optics and photonics technologies in a fundamental way. For example, light-based dissection methods like photo-isolation chemistry and photoselective sequencing use light to spatially select a region to profile, such as by triggering reverse transcription or removing a blocking group that prevents ligation of a sequencing adapter.
Spatial barcoding methods such as Visium and SeqScope deposit an ordered array of oligonucleotides on a glass side, each labeled with a barcode specific to each spatial position. A thin histological section is placed on top, with the cellular RNA diffusing to the barcoded oligonucleotides. Then, fluorescent complementary DNA produced through in situ reverse transcription is imaged with fluorescence microscopy, amplified and sequenced using traditional next-generation sequencing.
Although the field is still in its early days, research and development in spatial omics has already resulted in an abundance of technologies, each with its own advantages and disadvantages. Commercial “turnkey” systems—for instance, Visium by 10X Genomics, Vizgen MERSCOPE, and Akoya Biosciences’ PhenoCycler—have led to the wider adoption of methods previously too complex and costly for most labs to access.
“There’s a lot of excitement about these techniques, but like anything that gets a lot of buzz, it remains to be seen whether it’s going to keep building or crash down a little bit,” said Brian Beliveau, assistant professor of genome sciences at the University of Washington, USA. “The thing that researchers want to be mindful of, especially now that there are so many options out there, is what kinds of things spatial omics tools can do for them.”
Although the field is still in its early days, research and development in spatial omics has already resulted in an abundance of technologies, each with its own advantages and disadvantages.
Beliveau notes that, at this point, it may not be clear which tool to use in what scenario. Many spatial omics studies are more discovery-based—essentially, the researchers are going in without a hypothesis—which has value but perhaps less of an immediate impact on human health and disease.
Otherwise, he expects the field to make a push toward profiling thicker 3D volumes rather than slices of tissue a few micrometers thick. For image-based methods, a major challenge arises in terms of obtaining a sufficient signal-to-noise ratio during fluorescence microscopy, since biological tissue is a highly scattering medium. However, continued advances in optics and photonics, such as light-sheet imaging, have the potential to move spatial omics technologies closer to a true 3D realm.
“There are some questions where the context of an intact organ being sliced off, put into different sections, and analyzed separately just isn’t going to cut it,” he said. “So, trying to go volumetric is totally a top frontier, and it’s a real opportunity for the optics imaging community to make a contribution.”
In addition, Wählby believes that combining modalities—such as performing spatial transcriptomics and proteomics in parallel—as well as exploring other microscopic techniques could help advance the field. Her laboratory is interested in integrating second-harmonic imaging microscopy with in situ sequencing to understand what aspects of gene expression are causing the changes in cell and tissue structure associated with cancer progression.
“We need more information, but we also want to scale it up to more patient samples, so that we can get better statistics,” she said. “In this way, we can differentiate between patients who would benefit from different treatments, find biomarkers and get new ideas about how to improve treatments.”
Meeri Kim is a freelance science journalist based in Los Angeles, CA, USA.
For references and resources, visit: optica-opn.org/link/0424-omics.
