 [Enlarge image]A rapid, stain-free and cost-effective viral plaque detection system enabled by holographic imaging and deep learning can significantly reduce the detection time required by traditional viral plaque assays while eliminating chemical staining and manual PFU-counting procedures.
[Enlarge image]A rapid, stain-free and cost-effective viral plaque detection system enabled by holographic imaging and deep learning can significantly reduce the detection time required by traditional viral plaque assays while eliminating chemical staining and manual PFU-counting procedures.
In response to global health threats posed by viral infections, various virus detection and quantification techniques are used to accelerate the development of vaccines and antiviral drugs. Among these techniques, the viral plaque assay is the gold-standard method for quantifying the concentrations of replication-competent, cell-penetrating and actively reproducing viral particles (lytic virions) due to its unique ability to assess virus infectivity.
Traditional viral plaque assays, however, have several challenges. First, they are time-consuming, commonly requiring 2 to 14 days of incubation to observe plaque-forming units (PFUs) on cell samples. Second, a chemical staining process is needed to bring contrast to PFU samples to allow visual inspection by experts to manually count the resulting PFUs, an approach that is labor intensive and susceptible to human counting errors.
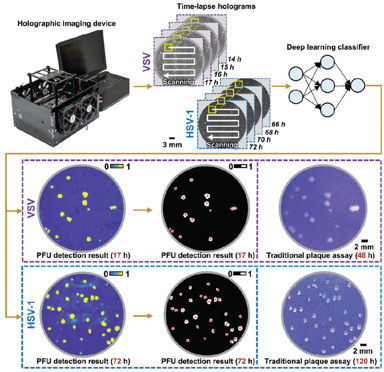
We have recently reported a rapid, stain-free and automated viral plaque detection system that uses time-lapse holographic imaging and deep learning.1 The system incorporates a high-throughput, cost-effective holographic imaging device to continuously monitor unstained virus-infected cells during the incubation process. At each imaging cycle, captured holograms that record the spatiotemporal features of the samples are reconstructed and processed by a deep-learning-based PFU classifier, outputting a PFU probability map with the locations and sizes of the detected PFUs.
We demonstrated the system’s efficacy using Vero E6 cells infected by different types of viruses, including vesicular stomatitis virus (VSV), herpes simplex virus (HSV-1), and encephalomyocarditis virus (EMCV). Using this system, the first cell-lysing events due to VSV replication were detected as early as 5 hours, and more than 90% of VSV viral plaques were detected with less than 20 hours of incubation—a savings of greater than 24 hours compared with the 48 hours required by traditional viral plaque assays. The system also reduced the detection time for HSV-1 PFUs by approximately 48 hours (from the 120 hours required for traditional viral plaque assays) and for EMCV PFUs by roughly 20 hours (from the 72 hours required for traditional assays). And we showed that this data-driven plaque assay can quantify PFUs over a tenfold-larger dynamic range of virus concentration than standard viral plaque assays.
Overall, the deep-learning-enabled holographic viral plaque detection system we demonstrated can significantly reduce the detection time required by traditional viral plaque assays and entirely eliminate chemical staining and manual counting procedures that lab technicians administer. This compact, low-cost, and automated PFU quantification device can be broadly used in virology research, vaccine development and clinical applications.
Researchers
Yuzhu Li, Tairan Liu, Hatice Ceylan Koydemir, Yijie Zhang, Ethan Yang, Merve Eryilmaz, Hongda Wang, Jingxi Li, Bijie Bai, Guangdong Ma and Aydogan Ozcan, University of California, Los Angeles, CA, USA
References
1. T. Liu, Y. Li et al. Nat. Biomed. Eng. 7, 1040 (2023).

